Forward, reverse, sense, antisense, first strand, second strand, unstranded. Different methods for sequencing RNA-Seq data can lead to differently stranded libraries all with different names. This can make it challenging to figure out how different kits compare to one another or what parameters to use with different software tools to make sure you are doing an analysis correctly. In this blog post, we will review common terms that are used for stranded libraries and common tool parameters for Eclipsebio kits.
DNA has its own stranded terminology that is separate than the terms used when we refer to RNA (A). The Watson strand, also called the + or top strand, is the strand of DNA that has its 5’ end in the short-arm telomere. The Crick strand, also called the – or bottom strand, is the strand of DNA that has its 3’ end in the long-arm telomere. This nomenclature is based off the physical orientation of the DNA strands and does not indicate if a given gene is in its sense or antisense orientation.
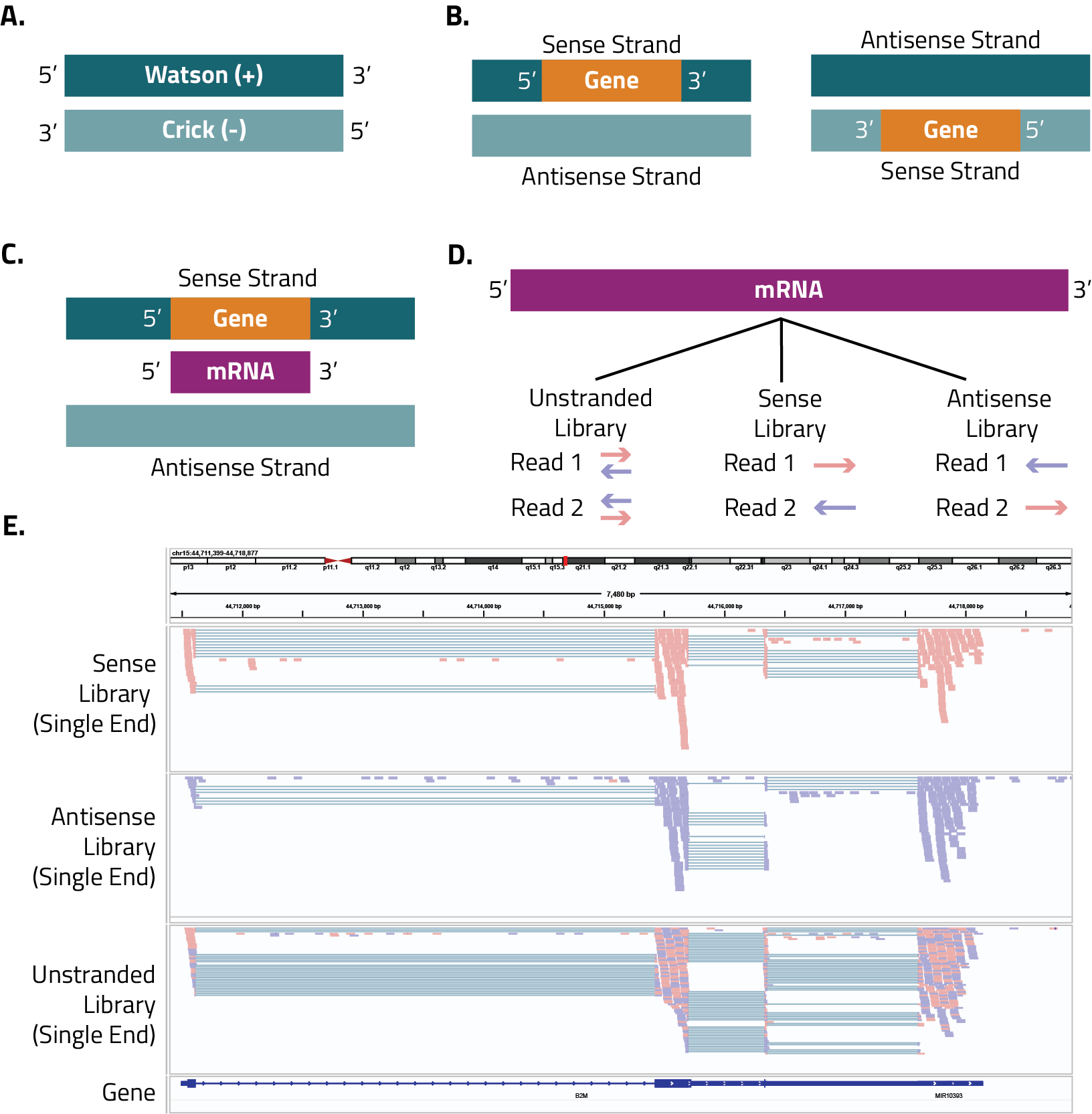
We can also refer to DNA strands in reference to individual protein coding genes (B). One strand is the “sense” strand, that is the strand that if read from right to left (5’ to 3’) carries the code for a protein. For this reason, the sense strand can also be called the coding strand, and can be on either the Watson or Crick strands. The sense labeling is based on the orientation of the gene and does not relate to the orientation of the DNA itself. The coding terminology is still used even if the gene is a noncoding sequence such as a lncRNA.
The antisense strand is the complementary strand to the sense strand, when it is read from right to left (5’ to 3’) its sequence cannot be translated into a protein and so it is sometimes called the noncoding strand. The antisense strand is used as the template for transcription (C), which means that the generated mRNA is in the sense direction (in other words, identical to the gene as coded in the sense DNA strand except thymines are now uracils).
In most RNA-Seq experiments, the mRNA is used as a template for two rounds of cDNA synthesis. One or both cDNA strands are sequenced and analyzed, and which strand gets sequenced depends on the library preparation method. Depending on the kit, these libraries can be considered either unstranded or directional (sense or antisense).
In an unstranded (non-stranded) preparation, both cDNA strands are sequenced. This means that we are not able to determine if the original RNA molecule had come from a sense mRNA sequence (same direction as the gene on the DNA) or an antisense noncoding RNA.
In a sense directional library (also called forward, stranded, same, first read, or second strand depending on the kit or software used used), the first read sequenced has the same orientation of the RNA molecule (D). If paired-end sequencing is performed, the second read will have the opposite orientation as the RNA molecule. These libraries are often made with ligation-based methods, and Eclipsebio kits are sense libraries.
| Library Type | Common Terms |
| Sense | Forward, stranded, same, first read, second strand |
| Antisense | Reverse, reverse stranded, second read, first strand |
In an antisense directional library (also called reverse, reverse stranded, second read, or first strand), the first read sequenced has the antisense orientation as the RNA molecule (D). If paired-end sequencing is performed, the second read will have the same orientation as the RNA molecule. These libraries are often made with dUTP-based methods.
Sometimes it can be difficult to determine the strandedness of the library after sequencing was performed. If that happens there are different tools available that can predict the strandedness of the library (RSeQC is one popular option). Alternatively, you can perform a simple alignment to your genome of interest and then view the reads on a genome browser and see if there is a bias to the reads. As shown in the figure (E), the unstranded library has a mixture of sense and antisense reads as shown by the equal mixtures of blue and red reads. The sense library has the majority of reads colored red, while the antisense library has the majority of reads colored blue. Importantly, you need to make this evaluation in the context of the gene. The example here is a gene that is going from 5’ to 3’ on the browser, if the gene was going the opposite direction the colors would be switched (sense library would be blue and antisense library would be red). Figuring out the strandedness of your library and the correct analysis parameters does not have to be a hassle.
Latest eBlogs
Charting a New Era: How RNA Is Unlocking N‑of‑1 Cures
RNA is unlocking the ability to develop life‑saving therapies at unprecedented speed. The recent success with baby KJ demonstrates how RNA medicine can move a personalized treatment from diagnosis to clinic in less than a year.
The three pillars of AI in RNA biology: why data is the hardest to get right
Artificial intelligence (AI) is transforming how we approach RNA research and drug discovery. In this eBlog we review how data is one of the key pillars for the successful use of AI.
