De-risk your RNA program from design to clinic.
Design, prototype, and characterize in an integrated loop. Every cycle generates empirical data that makes the next candidate better.
Get StartedFrom target to candidate.
Many RNA programs rely on standard approaches to sequence design and only discover dsRNA impurities, structural liabilities, or translation bottlenecks after manufacturing, when the cost to fix them is highest. eCOMPASS replaces that fragmented approach with a single partner that integrates AI-driven design, in-house prototyping, and sequencing-based characterization in a closed loop. You provide the amino acid sequence, and each cycle of empirical data produces a better candidate than the last.
Design
AI-driven RNA sequence optimization, balancing yield, expression, stability, and specificity to generate candidates tuned to your Target Product Profile.
Make
In-house manufacturing gets you from design to testable material fast. When you're ready to scale, we connect you to validated manufacturing partners.
Characterize
Nucleotide-level characterization from proprietary assays available nowhere else. Know not just what went wrong but why, from dsRNA sources to ribosome stalls.
Every lab-in-the-loop cycle provides a better candidate.
Design predicts. Validation confirms.
Even the most sophisticated sequence optimization produces candidates that should work, but dsRNA hotspots, ribosome stalls, and structural liabilities only emerge after synthesis. eCOMPASS closes that gap, feeding what characterization finds back into the next design cycle so each iteration is better by measurement, not just by prediction. That's the difference between a prediction and a decision you can defend.
Every cycle generates mechanistic insight, not just scores.
eCOMPASS generates over 150,000 data points per target amino acid sequence, covering identity, purity, potency, stability, and safety. These are some of the insights that drive candidate optimization and selection.
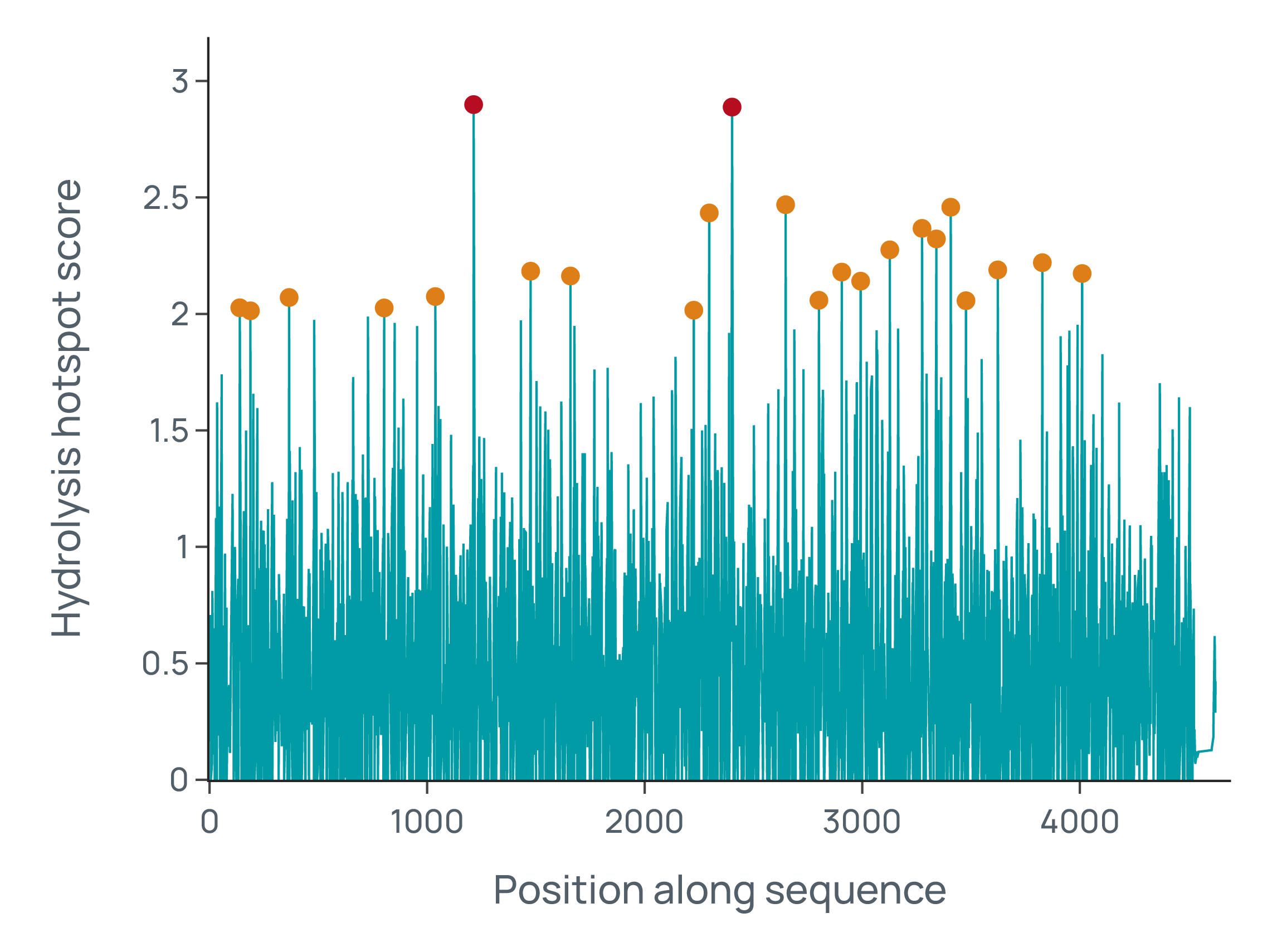
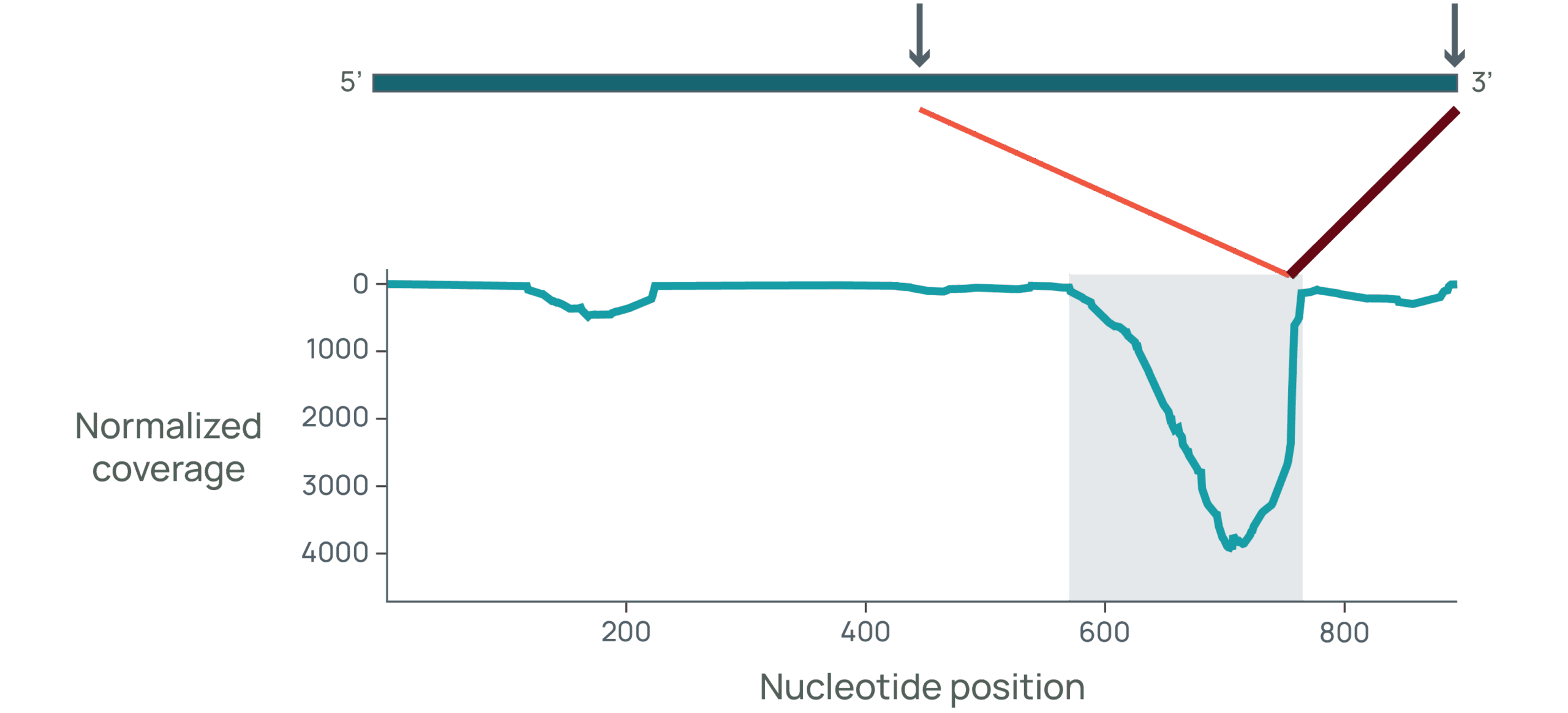
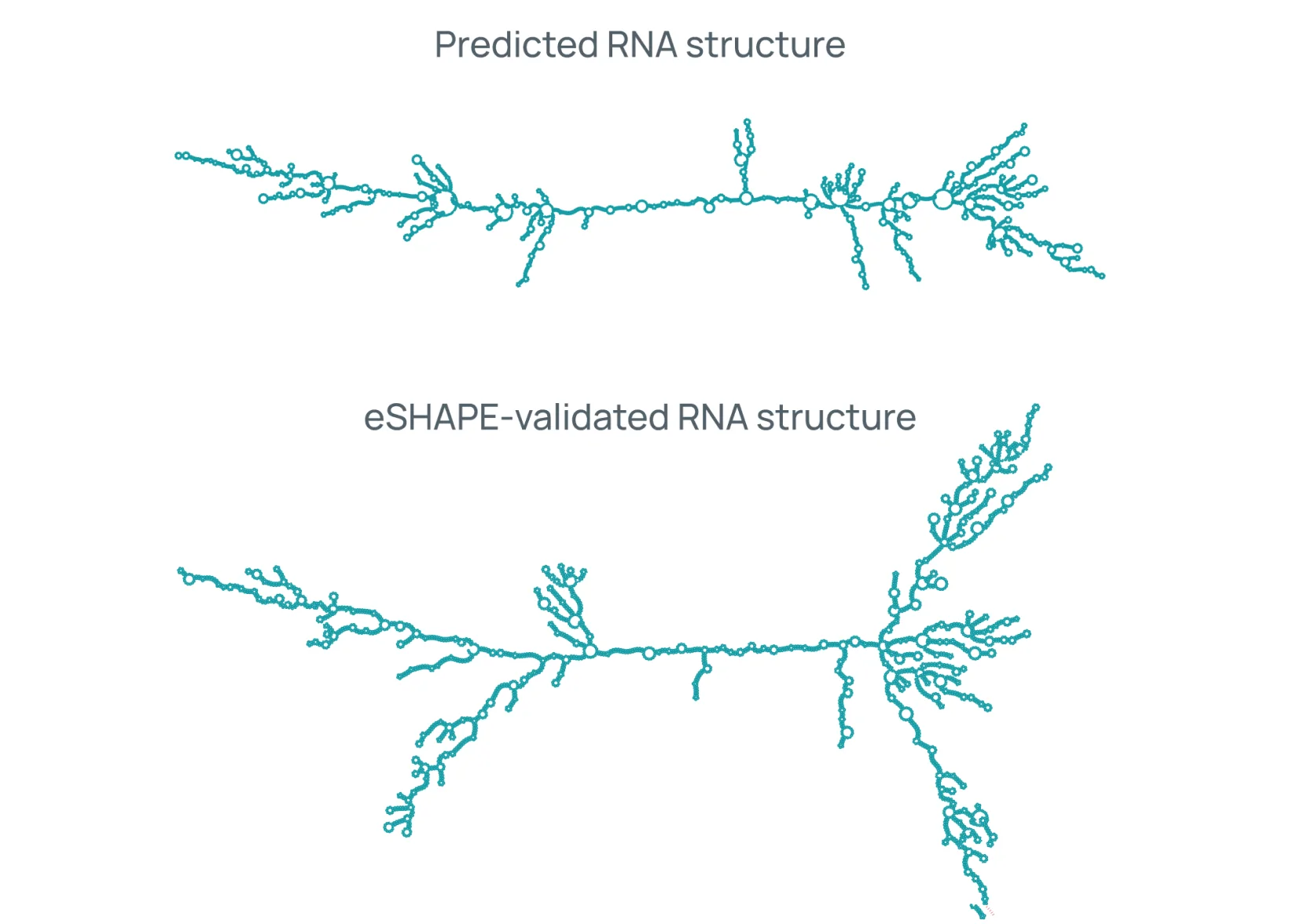
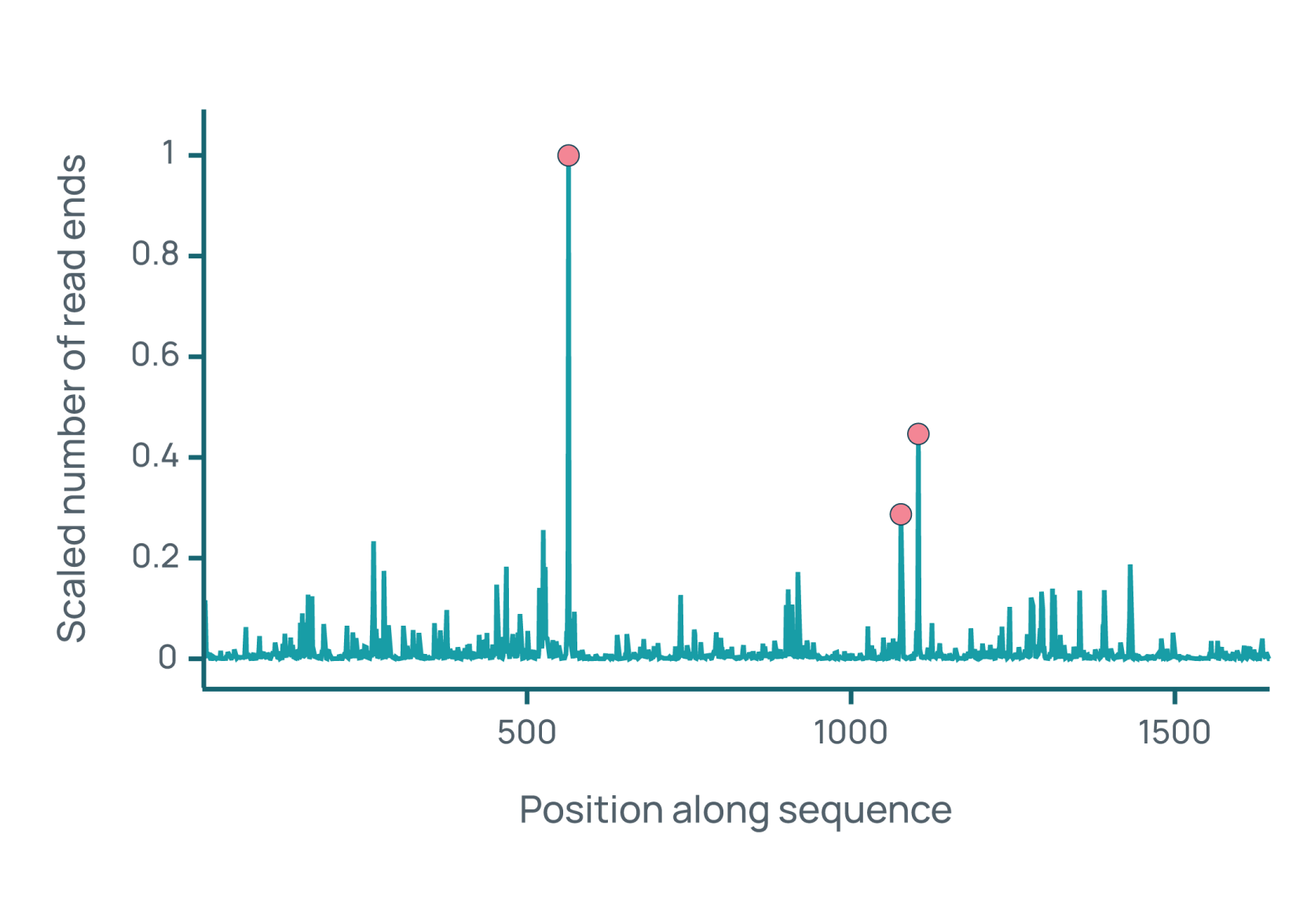
eCOMPASS in action.
This case study demonstrates eCOMPASS with the development of a gene editing therapy. We have also supported vaccine, CAR-T, and protein replacement programs for major biopharmas and biotechs.
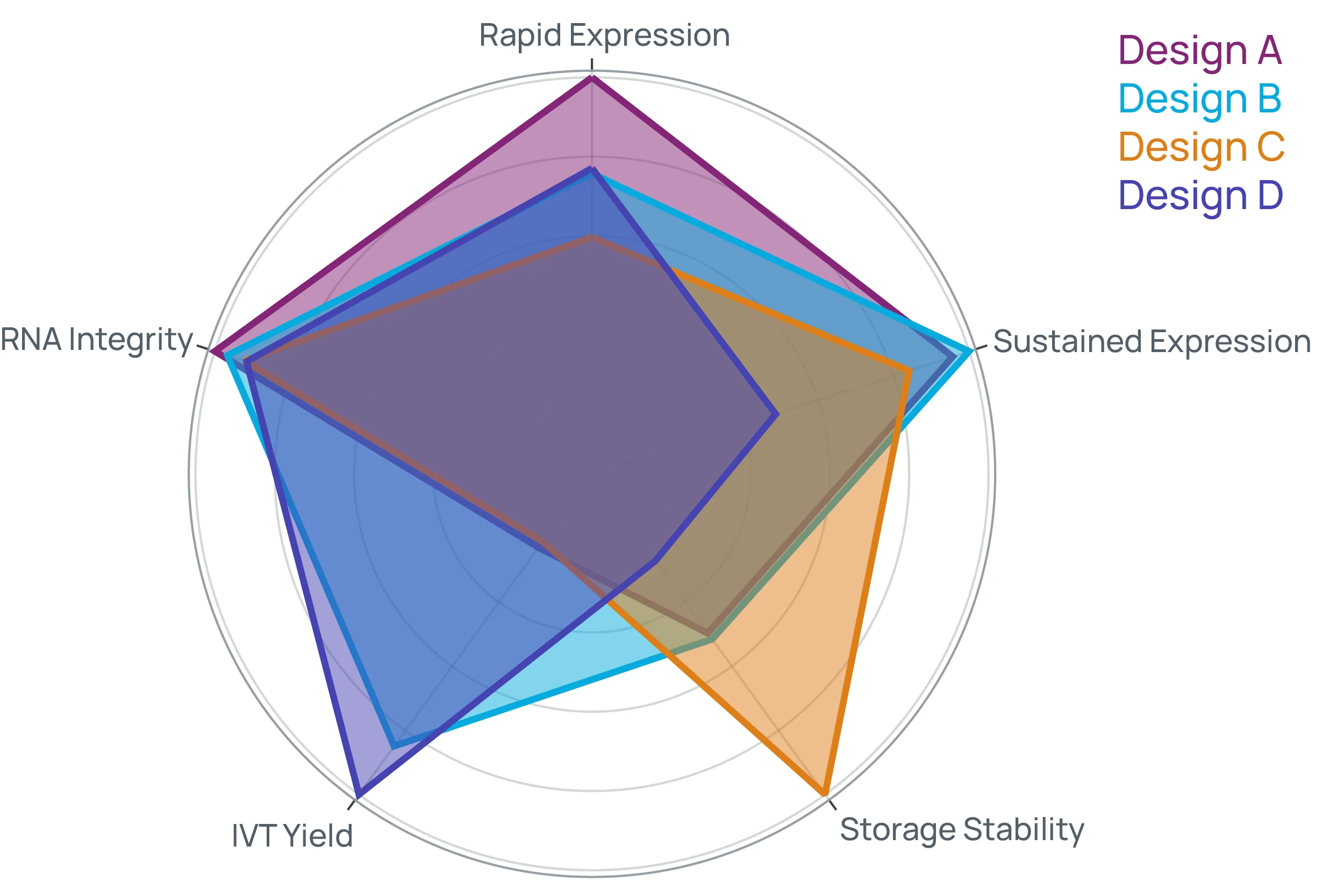
Multidimensional design outperforms single-metric optimization.
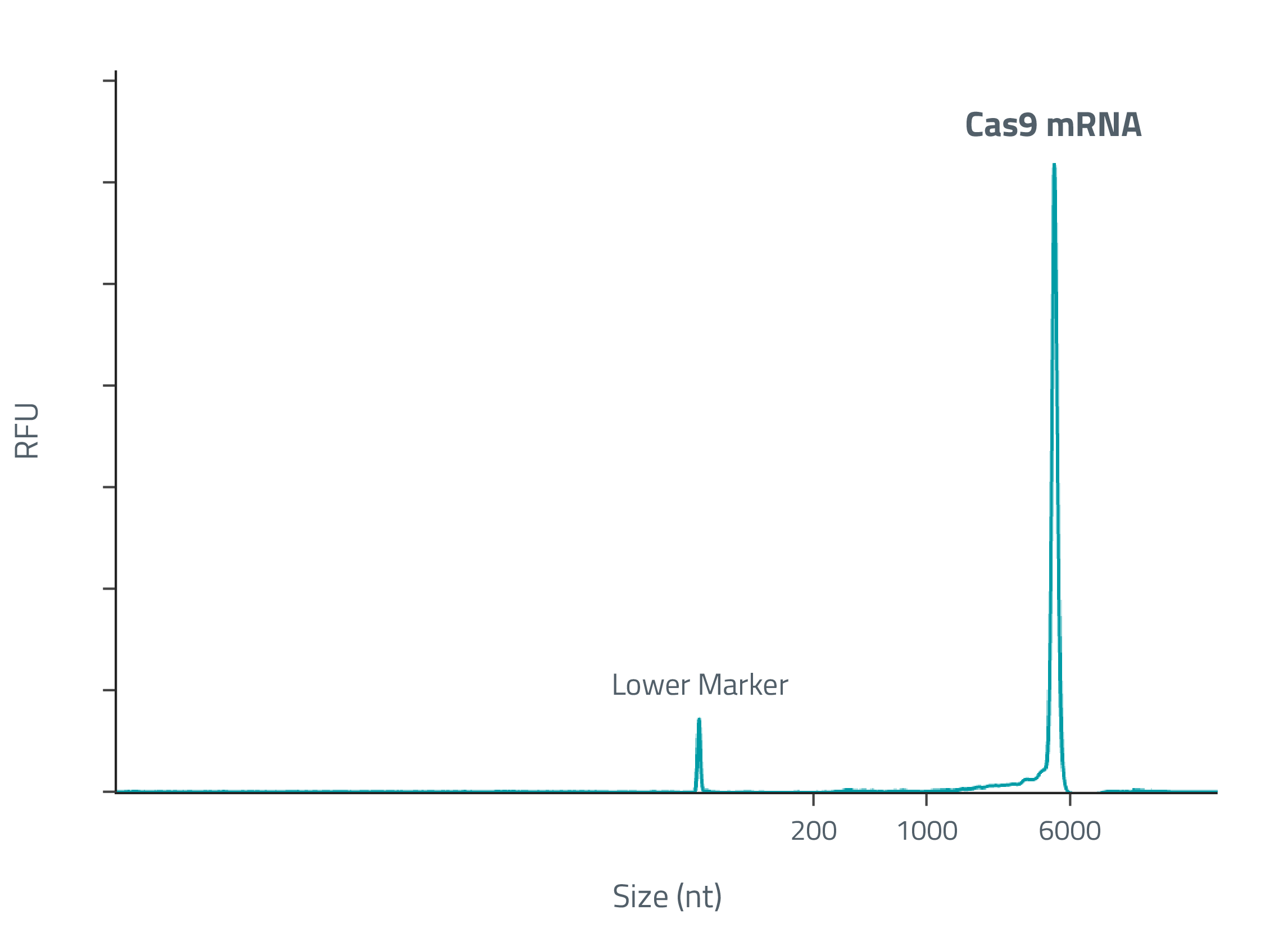
Cas9 mRNA sequences were optimized to balance yield, expression, and stability. Candidates were prototyped and characterized in-house. The resulting candidates outperformed both the baseline design and competing designs.
Our process.
You bring the program. We align on what success looks like, then iterate through design, prototyping, and characterization until your candidate is ready to advance. Most programs reach a characterized candidate within two to three cycles.
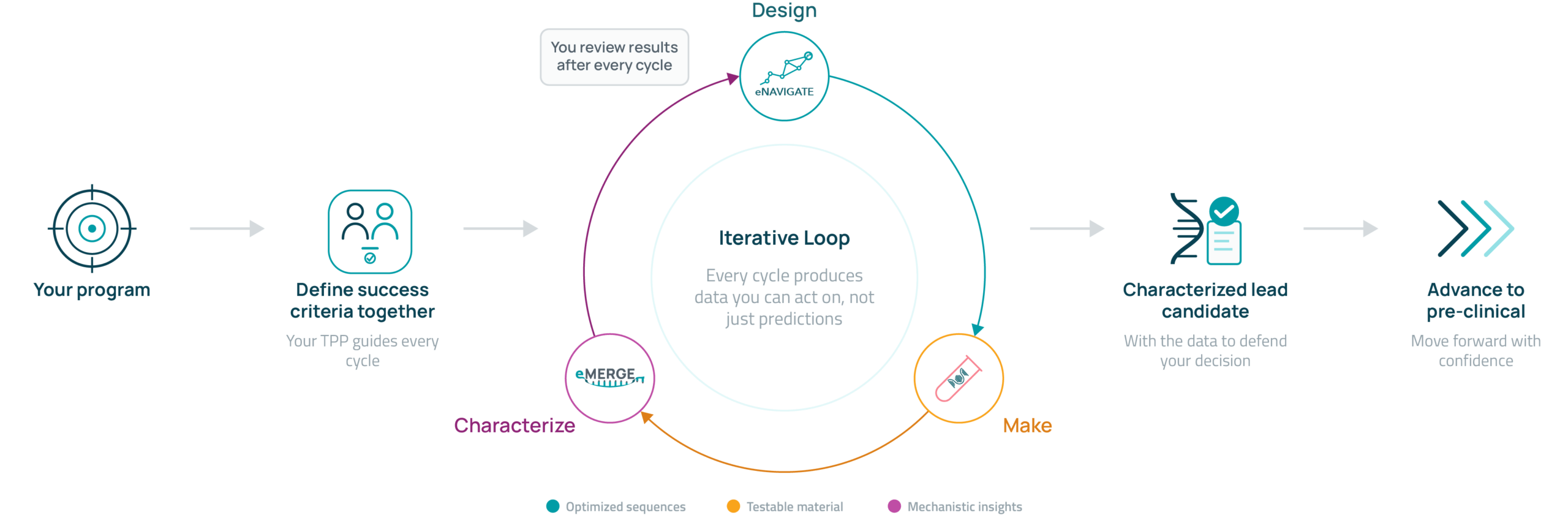
A closer look at each stage of eCOMPASS.
Trusted and proven.
Common questions about eCOMPASS.
What if we already use computational design tools?
What if we already have in-house characterization?
How is eCOMPASS different from working with a CDMO or an AI design platform?
What's the ROI on switching from our current approach?
How many cycles does a typical project take?
How do you handle IP and sequence confidentiality?
Ready to de-risk your RNA program?
We'll connect you with an RNA expert to explore how eCOMPASS fits your development goals.